Spatial atlas reveals how human organs begin to form after gastrulation
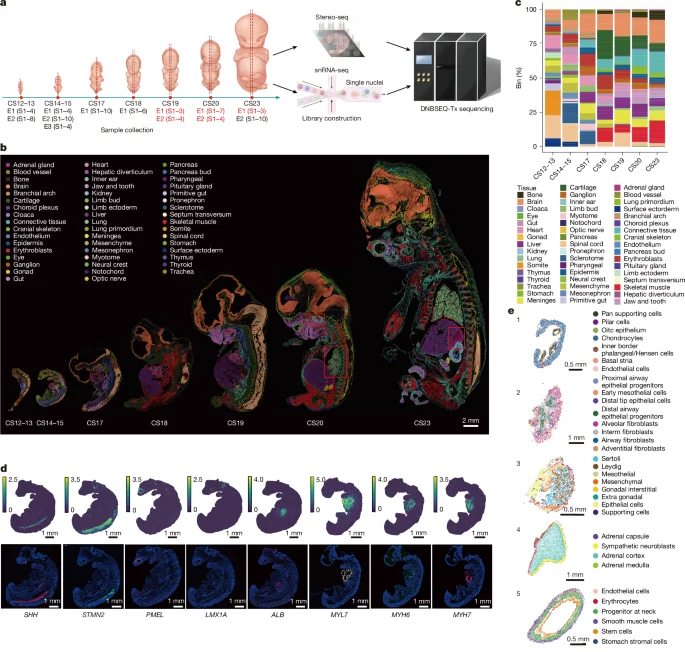
Pan et al. create a comprehensive spatiotemporal atlas of gene expression in 13 human embryos from Carnegie stage 12 to 23 using Stereo-seq and snRNA-seq, mapping 50 organs and 198 substructures, revealing context-specific regulatory networks and tissue identities, uncovering new gene functions in cardiac and brain development, and providing a public resource to visualize the embryonic transcriptional landscape and allelic expression dynamics across development.